Mathematics Its Content Methods And Meaning Pdf Archive
Welcome
A Structural View of Biology
This resource is powered by the Protein Data Bank archive-information about the 3D shapes of proteins, nucleic acids, and complex assemblies that helps students and researchers understand all aspects of biomedicine and agriculture, from protein synthesis to health and disease.
As a member of the wwPDB, the RCSB PDB curates and annotates PDB data.
The RCSB PDB builds upon the data by creating tools and resources for research and education in molecular biology, structural biology, computational biology, and beyond.

November Molecule of the Month

Acetohydroxyacid Synthase
Deposition Preparation Tools
Data Extraction
- pdb_extract: Extract and harvest data in PDBx/mmCIF format from structure determination programs
- SF-Tool: Convert structure factor files among various formats
Small Molecules
- Ligand Expo: Search the Chemical Component Dictionary for the IDs of released ligands
Data Format Conversion
- PDBML2CIF: Convert PDBML-format data into PDBx/mmCIF-format
- PointSuite: Generate symmetry records for macromolecular assemblies with point and helical symmetries
- MAXIT: Translate data between file formats and more
Validation Services
Validation reports contain an assessment of the quality of a structure and highlight specific concerns by considering the coordinates of the model, the experimental data and the fit between the two. Easily interpretable summary information that compares the quality of a model with that of other models in the archive will help users of PDB data to critically assess archived entries and to select the most appropriate structural models for their needs. These reports are developed using the recommendations of thewwPDB Validation Task Forces.
Reports for released entries are available from Structure Summary pages.
Validation reports for manuscript reviewers are created during annotation of deposited structures.
Information and example Validation Reports (at wwpdb.org).
Check your X-ray, NMR, or EM structures before depositing (standalone server).

Deposit 3D macromolecular structure data to the PDB
Have non-atomic coordinates, multi-scale structures obtained through integrative/hybrid (I/H) methods? Deposit at PDB-Dev which is a prototype deposition and archiving system for structural models obtained through integrative/hybrid (I/H) methods.
Questions about your deposition?

Basic Search
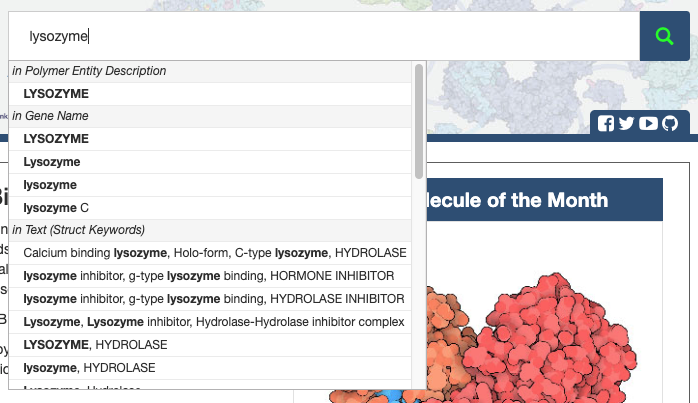
From any page on the site, a Basic Search can be run by entering a search term in the top Search Bar.
As you enter a term, you will see suggestions appearing in the dropdown menu that appears below the search bar. The suggestions are grouped by attribute name, indicating in the specific field or fields in which the search term was found.
You can click on an item in the dropdown menu to see results matching only that particular attribute, or click the
Search icon to see results matching multiple fields.
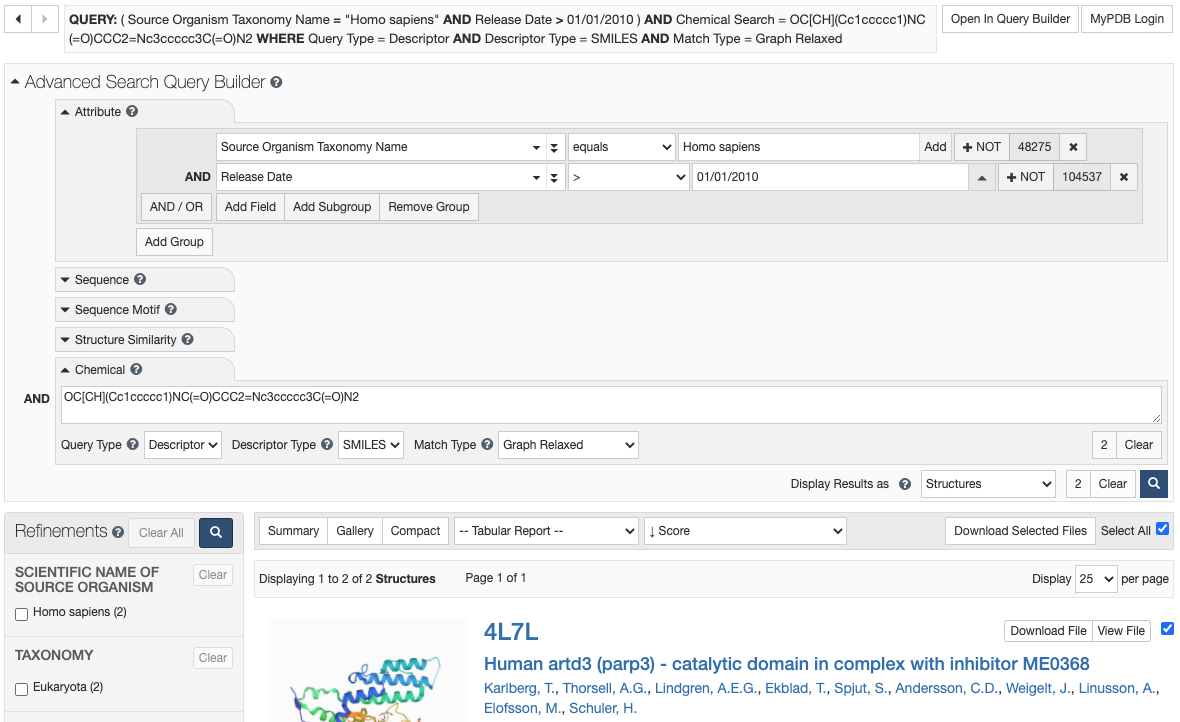
Advanced Search
The Advanced Search Query Builder tool allows you to construct complex boolean queries by specifying values for a wide range of structure attributes.
Text-based queries can be combined with sequence and structure similarity searches.
Search results can be returned at the structure, entity, or assembly level, and viewed in a variety of formats, for example, as a summary view, an images only gallery view, or in a Tabular Report format.
Any query and its results can be further refined by selecting additional criteria from the 'Refinements' panel.
Go to Advanced Search
Search by Sequences
Search protein and nucleic acid sequences using the mmseqs2 method to find similar protein or nucleic acid chains in the PDB.
The new Advanced Search Query Builder tool can be used to run sequence searches, and to combine the results with the other search criteria that are available.
Read Tutorial
Advanced Search - Sequence Search

Chemical Sketch Tool
You can search the PDB archive for a specific ligand or similar ligands based on the 2D chemical drawing of a molecule. In doing so you don't have to know the specific chemical descriptors (e.g., SMILES and/or InChI) because they will be automatically generated. You can also edit a molecule that is drawn or loaded into the tool to add or remove atoms or groups of atoms and then use the new molecule to query the PDB archive.

Browse by Annotation
PDB entries have been annotated by various ontologies and hierarchical classification schemes.
Search Ligands
Search ligands bound to macromolecules in the PDB by:
- SMILES String, InChI
- Chemical Formula
Ligand Search

Search by Drugs & Drug Targets
Drugs & Drug Targets in the PDB have been mapped to DrugBank. They have been made available through the integrated search system. You can find some search examples under the help examples page.
-

Drug-Target Complex: Atorvastatin bound to its target HMG-CoA reductase
-
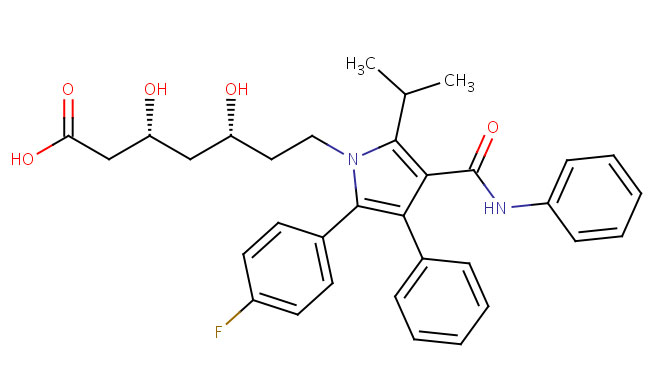
Drug: Atorvastatin (Lipitor)
Go to Advanced Search
Protein Feature View

Provides a graphical summary of biological and structural protein features of PDB entities and how they correspond to UniProtKB sequences. It loads features from RCSB PDB webservices and third party resources such as UniProtKB, CATH or SCOPe.
Learn more about Protein Feature View.
Examples: SARS-CoV-2 spike glycoprotein, H-Ras, and BRCA1
This feature is available in Structure Summary pages and Instance Sequence pages.
Genome View
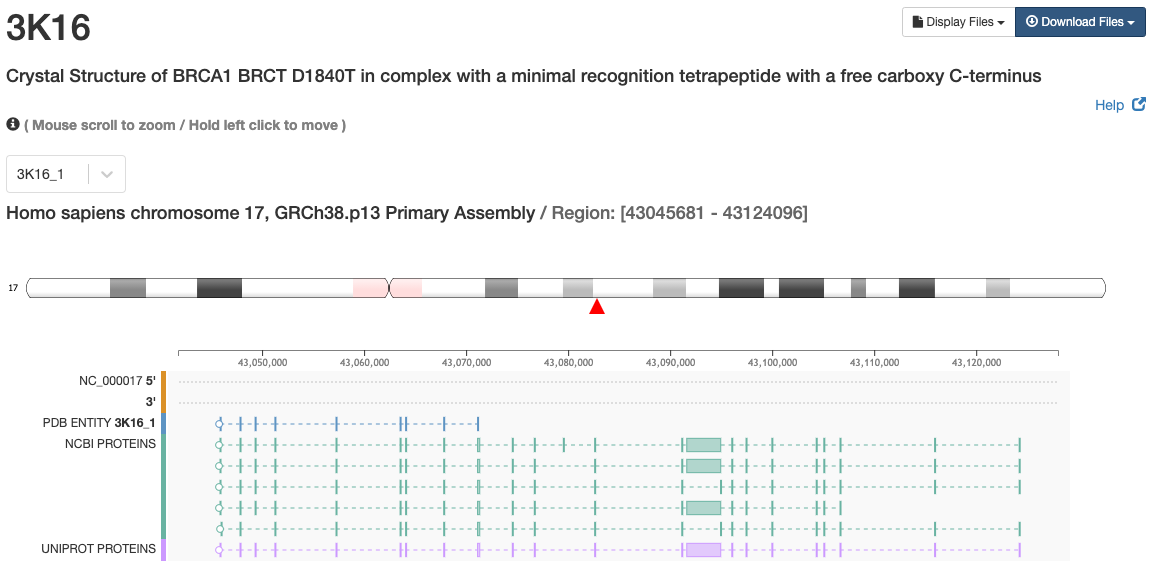
Illustrates the correspondences between PDB Entity sequences and genomes. When the relationships between PDB Entities and genes are available, PDB Entity sequences are mapped to their genome positions to show which regions of a gene are available in the Entity coordinates.
Learn more about Genome View.
Examples: Breast Cancer 1 (early onset) , SARS-CoV-2 spike glycoprotein and H-Ras (shown image).
This feature is available from the Structure Summary page Genome tab.

Pairwise Structure Alignment
The protein structure alignment tool allows for calculating pairwise structure alignments using different alignment methods. See the detailed documentation at this help page. Comparisons can be made for any protein in the PDB archive and for customized or local files not in the PDB.
Also, the standalone Mol* viewer application allows for calculating pairwise structure alignments based on sequence alignment. See detailed instructions on how to superpose structures at the Mol* help page.
Protein Symmetry
The Mol* symmetry display mode (select the Assembly Symmetry button) highlights global, local, and helical symmetry among subunits. The view displays the symmetry axes, a polyhedron that reflects the symmetry, and a color scheme that emphasizes the symmetry.
-
Hemoglobin

PDB ID:4HHB
-
Streptavidin
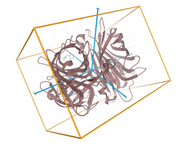
PDB ID:1STP
-
Inovirus

PDB ID:1IFD
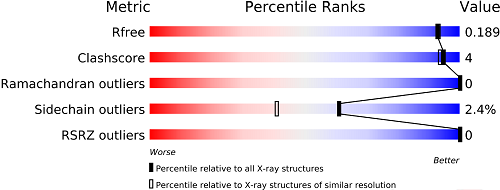
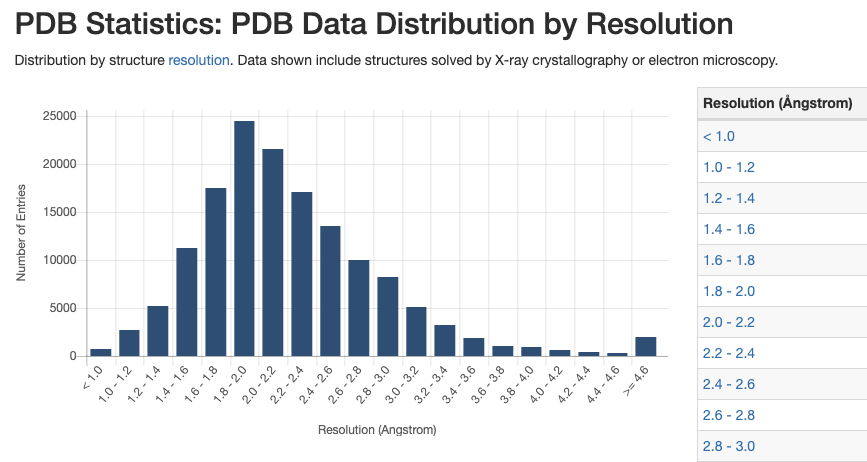
Structure Quality
Structure Summary pages provide access to information about structure quality.
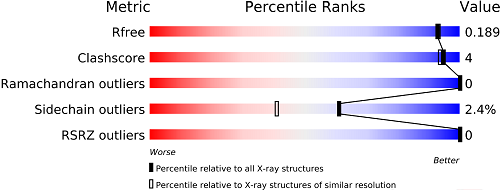
The slider graphic compares important global quality indicators for a given structure with the PDB archive. Global percentile ranks (black vertical boxes) are calculated with respect to all X-ray structures available prior to 2011. Resolution-specific percentile ranks (white vertical boxes) are calculated considering entries with similar resolution.
This graphic is from the wwPDBValidation Report , which provides a more detailed assessment of the quality of a structure and highlights specific concerns. These reports were created using the recommendations of wwPDB Validation Task Forces.The full wwPDB Validation Report PDF is available for download.PDFs ofRamachandran plots (created by MolProbity) are provided to offer an independent method to evaluate the conformational quality of protein structures.
For examples, view the Structure Summary pages for 1CBS (1.8Å structure of a small protein and a ligand, an entry with better overall quality relative to all X-ray structures) and 1FCC (a 3.2Å structure with worse overall quality relative to all X-ray structures).

Map Genomic Locations to/from PDB
The 1D-coordinates API provides means to map between genomic coordinates and PDB positions. Find more information about this and other APIs in the webservices page.
1D-coordinates API tutorial
EPPIC Biological Assemblies
EPPIC (Evolutionary Protein-Protein Interface Classifier) provides value-added information about biological assemblies in the PDB. This web server classifies interfaces present in protein crystals to distinguish biological interfaces from crystal contacts. EPPIC Version 3 enumerates all possible symmetric assemblies with a prediction of the most likely assembly based on probabilistic scores from pairwise evolutionary scoring.

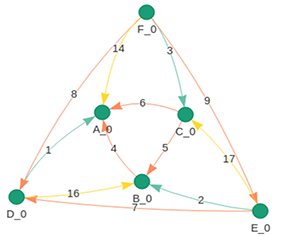
PDB Citation MeSH Network Explorer
The National Library of Medicine assigns MeSH (Medical Subject Headings) from a controlled vocabulary to index articles for PubMed. MeSH terms typically appear in a hierarchical tree structure that starts with 16 main branches. The PDB Citation MeSH Network Explorer flattens these trees into co-occurence networks of MeSH terms associated with PDB entries. Each node on the graph is a publication, and nodes are linked when they share MeSH terms.
Download Coordinate & Experimental Data Files
By entering PDB IDs, multiple files can be downloaded in batches containg one or more file formats.
Coordinate Data Files can be downloaded in the following formats:
- PDB
- PDBx/mmCIF
- PDBML/XML
- PDBML/XML (Header only)
- Biological Assemblies in PDB
- Biological Assemblies in PDBx/mmCIF
Experimental Data Files can be downloaded in the following formats:
- Structure Factors
- NMR Restraints
- Chemical Shifts
- NMR Restraints v2
Go to the Downloads Page
Download: Sequences
By entering PDB IDs, sequences can be downloaded in FASTA format.
Sequences can be provided for any of these identifiers:
- Entry IDs
- Entity IDs
- Asym IDs
Go to the Sequences Downloads Page
Download: Ligands
By entering chemical component IDs, SDF files with ligand coordinates can be downloaded.
Downloads are provided for:
- Coordinates of first chemical component instance from each PDB entry
- Coordinates of all chemical component instances from each PDB entry
- Ideal coordinates from Chemical Component Dictionary
Go to the Ligands Downloads Page
Molecular explorations
through biology and medicine
PDB-101 is an online portal for teachers, students, and the general public to promote exploration in the world of proteins and nucleic acids.
Browse all PDB-101 resources by biological theme or start exploring:
-
Molecule of the Month
Presents short accounts on selected molecules from the Protein Data Bank.
-

News and Events
Upcoming meetings and events RCSB will hold
-

Educational Resources
Access materials that promote exploration in the world of proteins and nucleic acids.
-

Guide to PDB Data
Understanding PDB Data is a reference to help explore and interpret individual PDB entries.
-
Curricula
Authentic, hands-on teaching materials, individual and group activities.
-
Geis Digital Archive
View iconic illustrations by the gifted artist Irving Geis (1908-1997) in context with PDB structures and educational information.
Mathematics Its Content Methods And Meaning Pdf Archive
Source: https://www.rcsb.org/
Posted by: wickerbutfult1935.blogspot.com

0 Response to "Mathematics Its Content Methods And Meaning Pdf Archive"
Post a Comment